In April 2026, the Dutch-flagged polar expedition cruise ship MS Hondius departed from Ushuaia, Argentina. On 6 April, a passenger developed fever and headache, and died on board five days later. His wife also fell ill and died after being transferred to South Africa. Subsequently, a 69-year-old British passenger tested positive for hantavirus and remains in intensive care. As of 13 May, the World Health Organization reported 11 cases and 3 deaths associated with the outbreak on the ship.
I. Virus Overview and Pathogenic Types
Hantaviruses are a group of RNA viruses carried mainly by rodents, belonging to the genus Orthohantavirus in the family Bunyaviridae. Their genome consists of three segments (L, M, and S) , encoding RNA-dependent RNA polymerase, glycoproteins, and nucleocapsid protein, respectively. Based on geographic distribution and clinical manifestations, hantaviruses are classified into two major groups:
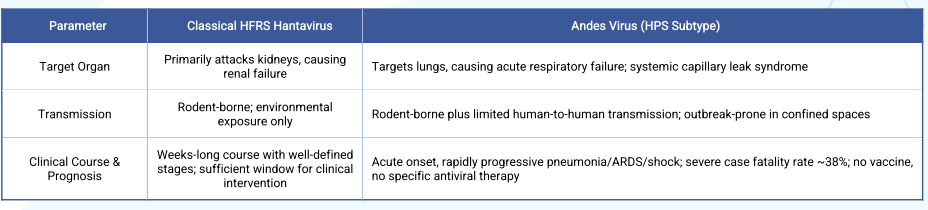
Old World hantaviruses (found in Europe, Asia, and Africa) cause Haemorrhagic Fever with Renal Syndrome (HFRS) . Representative viruses include Hantaan virus (HTNV), Seoul virus (SEOV), and Puumala virus (PUUV). They primarily target the kidneys, leading to renal impairment, haemorrhagic manifestations, and in severe cases, shock. Case fatality rates vary by virus type but generally range from <1% to 5-15% .
New World hantaviruses (circulating in North and South America) cause Hantavirus Pulmonary Syndrome (HPS) . Representative viruses include Sin Nombre virus (SNV) and Andes virus (ANDV). They primarily target the lungs, causing rapid-onset non-cardiogenic pulmonary oedema, severe respiratory distress, and cardiovascular compromise. Initial symptoms are flu-like (fever, myalgia, gastrointestinal complaints), but the disease can quickly progress to respiratory failure, with case fatality rates reaching 30-50% .
The cruise ship outbreak was mainly caused by Andes virus (ANDV) , which belongs to the Hantavirus Pulmonary Syndrome (HPS) subtype.
II. Traditional HFRS Associated Hantaviruses vs. Andes Virus (HPS)
III. Modes of Transmission
Hantavirus is mainly transmitted through direct contact with the faeces, saliva, or urine of infected rodents, or by inhalation of aerosolised excreta. Other possible routes include:
-Being bitten or scratched by infected rodents;
-Eating food contaminated with infected rodent urine, droppings, or saliva;
-Touching the eyes, nose, or mouth after contacting contaminated articles.
Human-to-human transmission is extremely rare, but has been confirmed for Andes virus (ANDV).
IV. Global Hantavirus Contact Tracing – Nucleic Acid Testing and Sequencing as Core Technologies
Following the ANDV outbreak, health authorities in more than ten countries – including Spain, the United Kingdom, the United States, Canada, and Switzerland – simultaneously launched contact tracing and quarantine monitoring. France, the United States, and others have continued to track the health status of their citizens who returned from the affected ship.
In this multi-country emergency response, nucleic acid testing and genomic sequencing have played an indispensable core role:
-Nucleic acid testing provides definitive evidence for rapid pathogen identification;
-Genomic sequencing offers irreplaceable molecular evidence for tracing the source of infection, confirming transmission chains, and assessing viral changes.
V. Detection Solutions for Hantavirus (Especially HPS Type)
1. Early Pathogen Identification
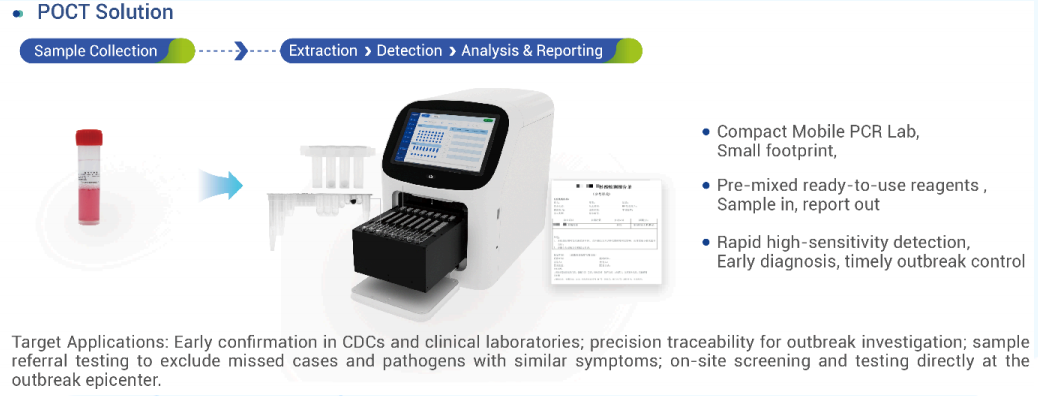
1.1 Point-of-Care Testing (POCT)
Precise screening and emergency testing – sample-to-result in 1 hour.

1.2 Conventional PCR

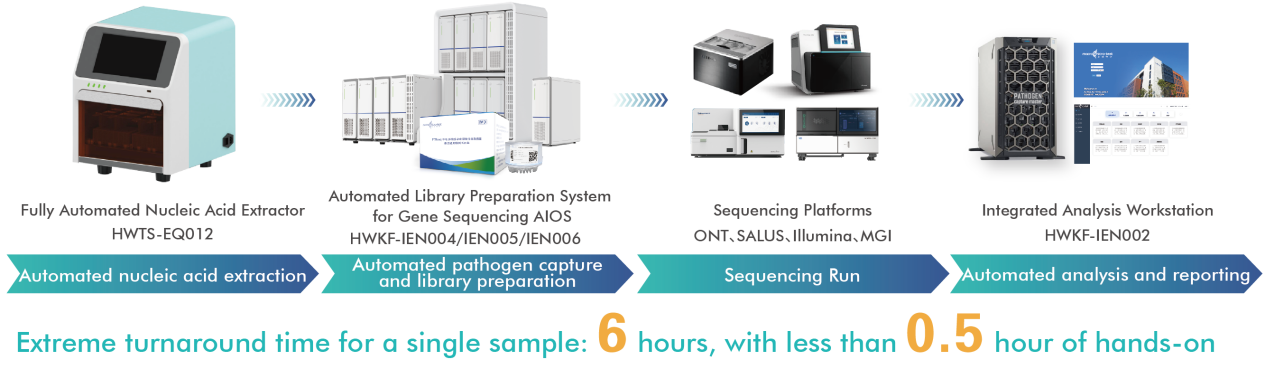
2. Sequencing Solutions
For pathogen identification and outbreak tracing in cases of fever of unknown origin, respiratory distress, or haemorrhagic manifestations.

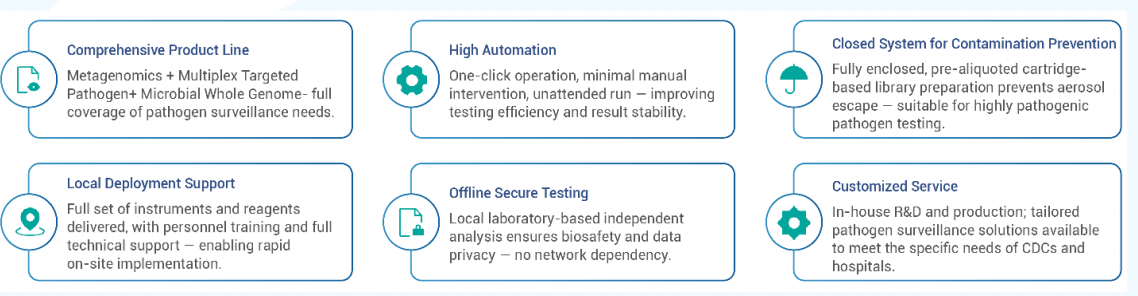
Core features:
VI. Verified Performance Data
Probe capture-based whole-genome sequencing of hantavirus-positive samples achieved >99% genome coverage at 30× depth.
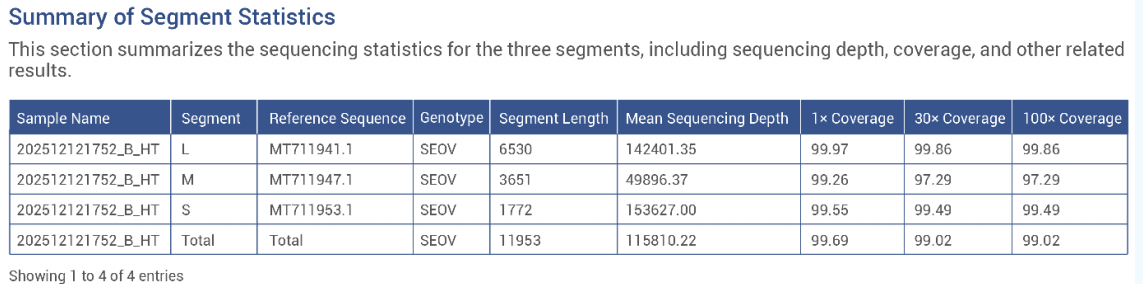
Wastewater-based hantavirus genotyping and subtype proportion analysis is feasible.
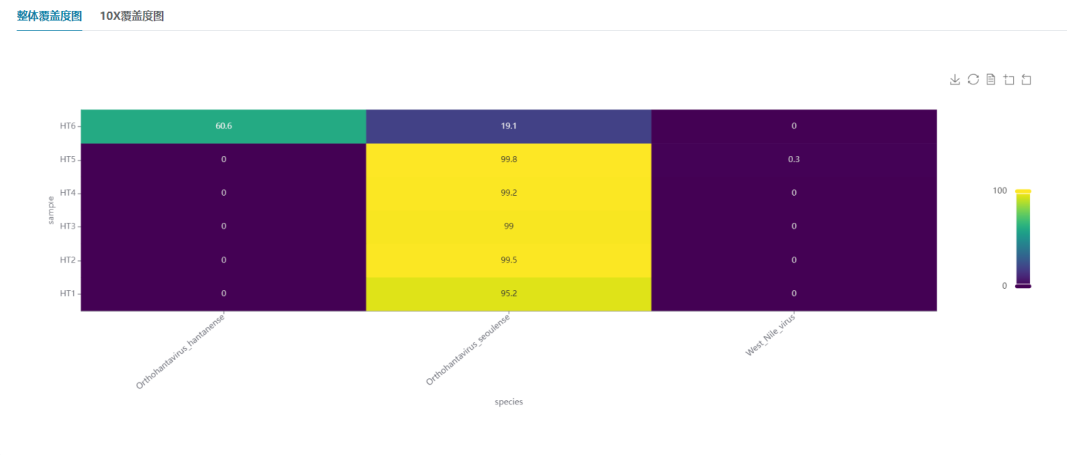
VII. Related Kits
Post time: May-14-2026