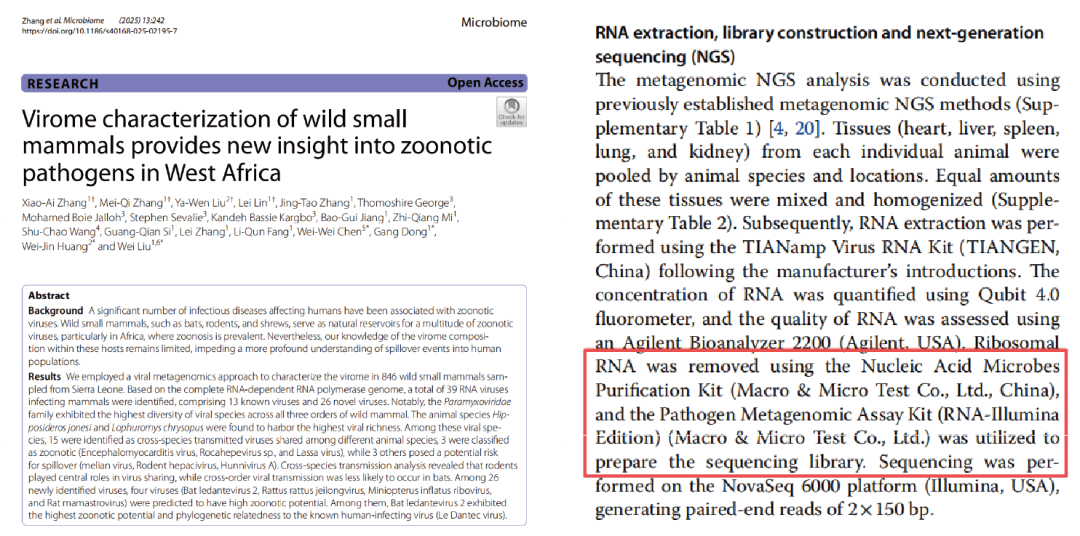
A recent study published in Microbiome conducted viral metagenomic analysis on 846 wild small mammals—including bats, rodents, and shrews—collected in Sierra Leone, West Africa. The study identified a total of 39 mammal-associated RNA viruses, comprising 26 novel and 13 previously known viruses. Among these, the Paramyxoviridae family exhibited the highest diversity, while rodents harbored the greatest number of viral species (n = 26).
Zoonotic risk assessment revealed three known zoonotic viruses—encephalomyocarditis virus, Lassa virus, and Rocahepevirus sp.—as well as three viruses with potential spillover risk: Melian virus, rodent hepatitis virus, and Hunnivirus A. Notably, among the newly identified viruses, Bat ledantevirus 2 showed the closest phylogenetic relationship to the human-infecting Le Dantec virus. Serological analysis further detected neutralizing antibodies against this virus in 2.8% of local residents, suggesting prior, likely undetected, human exposure.
These findings highlight the presence of a substantial rodent-dominated viral reservoir in West Africa and underscore the critical importance of integrated surveillance strategies at the human–animal interface. Combining metagenomic screening with serological validation provides a robust framework for identifying viruses with zoonotic and spillover potential.

Over the past decade, more than 60% of emerging infectious diseases in humans have originated from animal reservoirs, with bats, rodents, and shrews recognized as key hosts of zoonotic viruses. Africa is widely regarded as a hotspot for zoonotic diseases. For instance, Sierra Leone reported over 28,000 cases during the 2014–2016 Ebola outbreak.
Despite the significant burden of zoonotic diseases in this region, the diversity and distribution of viruses in wild small mammals remain insufficiently characterized. To address this gap, researchers conducted a systematic virome analysis of 846 wild small mammals captured across three sites in Sierra Leone between 2018 and 2023. The study aimed to characterize viral diversity, identify candidates with cross-species transmission potential, assess zoonotic risk, and generate evidence to support early warning systems for emerging infectious diseases.
Core Methods
The study applied a comprehensive viral metagenomics workflow:
- Sample processing: Heart, liver, spleen, lung, and kidney tissues were collected, pooled, homogenized, and subjected to total RNA extraction.
- Sequencing and assembly: Ribosomal RNA depletion was performed prior to library construction, followed by high-throughput sequencing using the Illumina NovaSeq 6000 platform. Viral contigs were assembled de novo.
- Virus identification: Viruses were identified based on RNA-dependent RNA polymerase (RdRp) gene alignment. Only vertebrate-associated viruses were retained, excluding bacterial, fungal, and plant viruses.
- Bioinformatics analysis: Phylogenetic reconstruction, recombination analysis, cross-species transmission network modeling, and zoonotic risk assessment were conducted.
- Serological validation: A VSV-based pseudovirus neutralization assay was developed for Bat ledantevirus 2. Neutralizing antibodies were detected in 2.8% of human sera, providing evidence of potential zoonotic transmission.
Study Results
1. Viral Discovery and Diversity
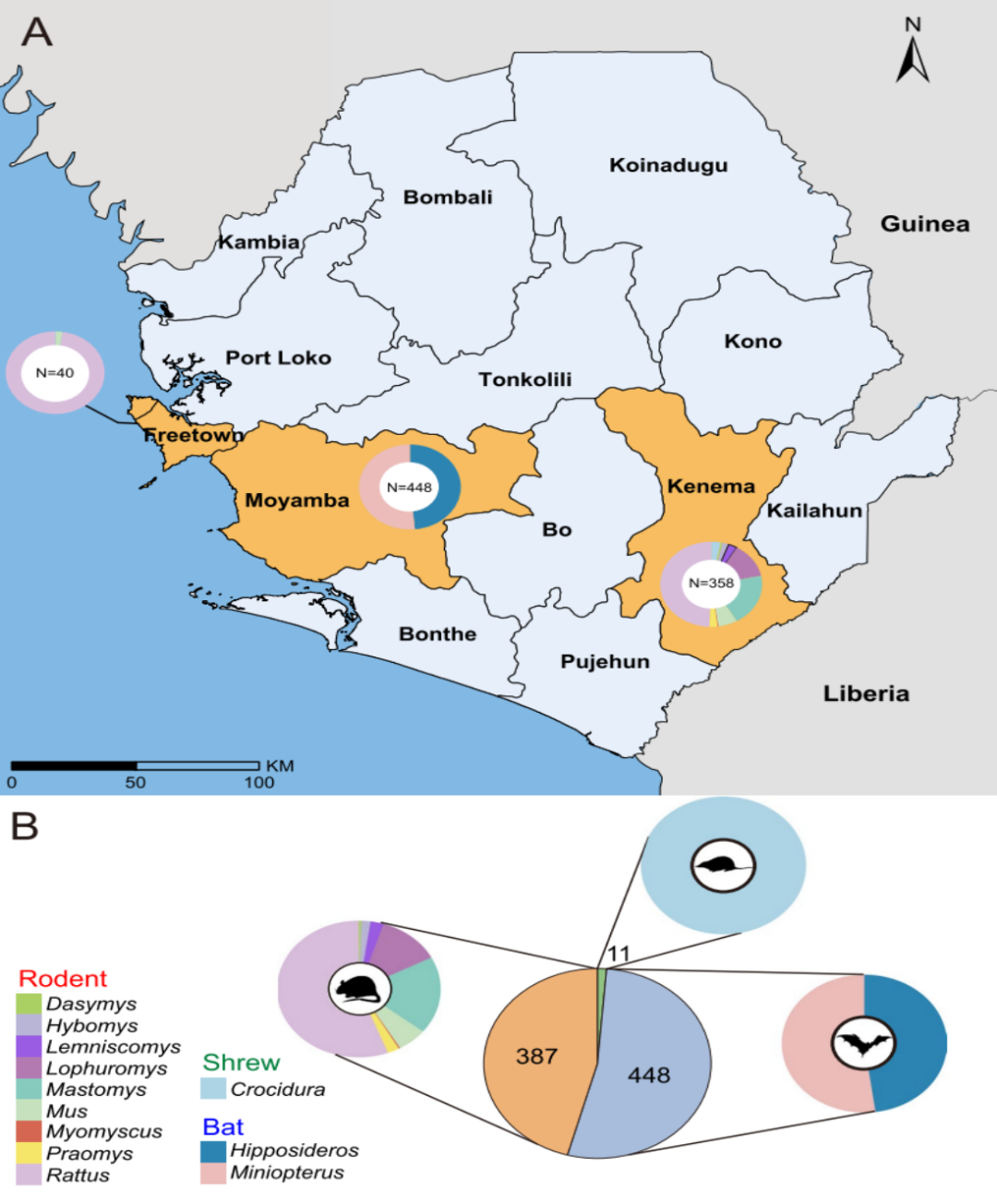
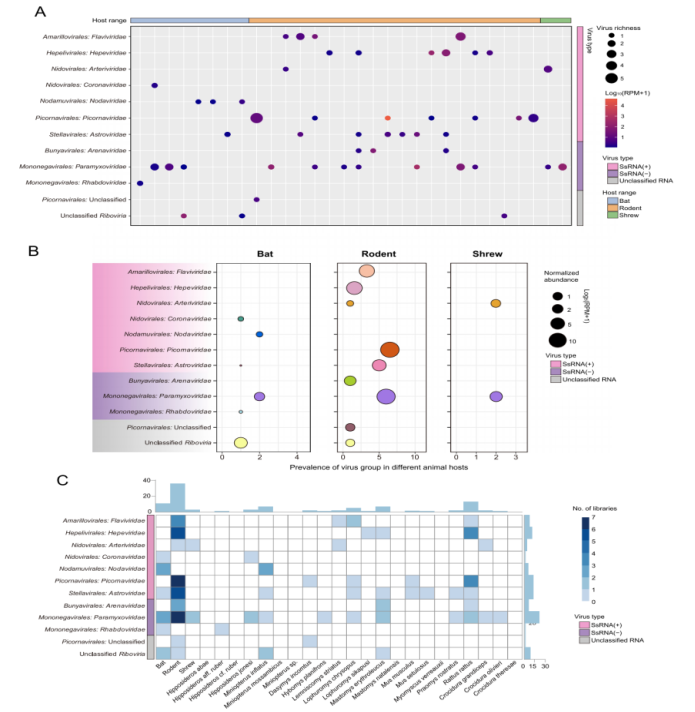
This study conducted transcriptomic sequencing analysis on 846 wild animals collected in Sierra Leone, including rodents, bats, and shrews. Based on complete RNA-dependent RNA polymerase (RdRp) gene sequences, a total of 39 mammal-associated RNA viruses were identified, comprising 13 previously known viruses and 26 novel viruses.
In terms of viral composition, the Paramyxoviridae family exhibited the highest level of diversity across all three host orders, followed by Astroviridae and Picornaviridae. Regarding host distribution, rodents contributed the greatest viral diversity, harboring a total of 26 virus species, indicating their prominent role as reservoirs of viral diversity in the region.
2. Zoonotic Risk
Zoonotic risk assessment identified three known zoonotic viruses: encephalomyocarditis virus, Lassa virus, and Rocahepevirus species. In addition, three viruses—Melian virus, rodent hepatitis virus, and Hunnivirus A—were identified as having potential spillover risk.
Among the 26 newly discovered viruses, four were predicted to possess high zoonotic potential based on phylogenetic and genomic characteristics. Notably, Bat ledantevirus 2 showed the closest phylogenetic relationship to the known human-infecting Le Dantec virus.
Subsequent serological investigation further supported this finding, as neutralizing antibodies against Bat ledantevirus 2 were detected in 2.8% of sera from local residents. This result suggests that unrecognized or asymptomatic infections may have already occurred within the human population, highlighting a potential but previously undetected zoonotic transmission pathway.
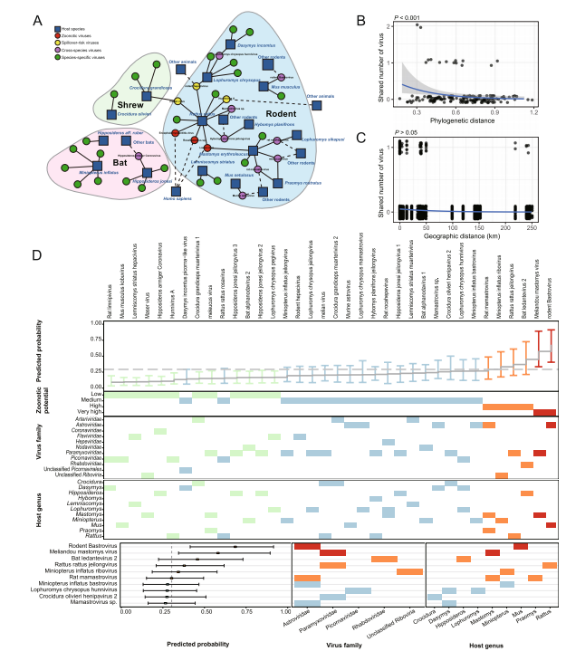
3. Cross-Species Transmission Dynamics
Cross-species transmission analysis demonstrated that rodents occupy a central position within the viral sharing network, acting as key nodes that facilitate viral exchange between host species. A total of 15 viruses were identified as having the potential for cross-species transmission.
Further analysis of cross-order transmission patterns indicated that viral sharing occurred more frequently among hosts within the same taxonomic order, suggesting that host relatedness plays an important role in transmission dynamics. In contrast, bats exhibited a relatively lower capacity for cross-order transmission.
Importantly, evidence of host range expansion was observed in certain viruses. For example, Melian virus, which had previously been considered specific to shrews, was also detected in rodents in this study, indicating a potential shift in host adaptability and an increased risk of broader transmission.
Conclusions and Public Health Implications
- High virome diversity in wild small mammals: The discovery of 39 RNA viruses, including 26 novel species, reveals a large virus reservoir in the region and reports for the first time novel viruses with high zoonotic potential (e.g., Bat ledantevirus 2).
- Rodents as priority surveillance targets: Rodents act as key hubs for viral transmission and carry the highest viral diversity, representing the greatest spillover risk.
- Need for integrated surveillance strategies: The findings support prioritizing rodents in active surveillance programs and implementing integrated approaches combining metagenomics, serology, and ecological monitoring at human–wildlife interfaces.
Overall, this study provides critical evidence to support early warning systems and risk assessment frameworks for emerging zoonotic diseases, reinforcing the importance of proactive surveillance in high-risk regions.
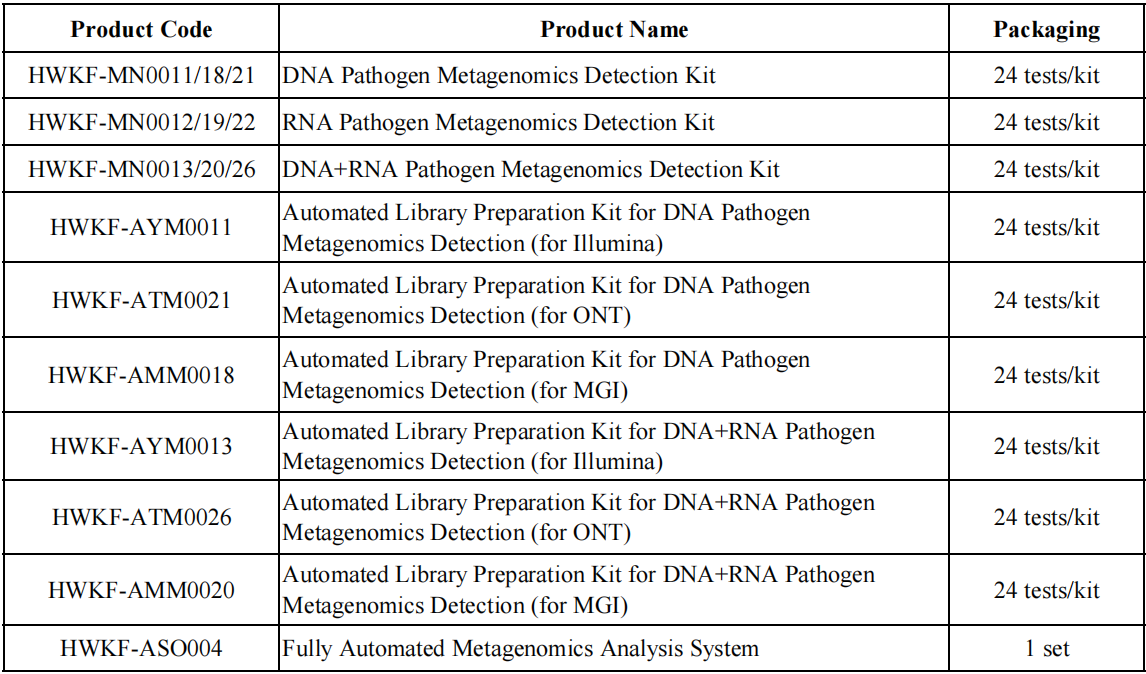
Product Information
Post time: Mar-23-2026